Data Structures and Algorithms: A Bioinformatics Analysis
VerifiedAdded on 2023/06/16
|9
|1285
|294
Homework Assignment
AI Summary
This assignment delves into various aspects of data structures and algorithms within the field of bioinformatics. It includes constructing keyword and suffix trees from a given DNA sequence, calculating the number of nucleotide sequences for a protein sequence, describing a pseudocode for identifying informative sites in a Multiple Sequence Alignment (MSA), explaining gene finding algorithms, detailing the BLAST algorithm and its statistical measures, and analyzing an R script for statistical analysis, including modifications to display a specified number of centroids. The solutions provided cover the processes and methodologies used in bioinformatics to analyze biological data and extract meaningful insights.

Running Head: DATA STRUCTURES AND ALGORITHMS 1
Data Structures and Algorithms
[Name of Student]
[Instructional Affiliation]
[Date of Submission]
Data Structures and Algorithms
[Name of Student]
[Instructional Affiliation]
[Date of Submission]
Paraphrase This Document
Need a fresh take? Get an instant paraphrase of this document with our AI Paraphraser
Data Structures and Algorithms 2
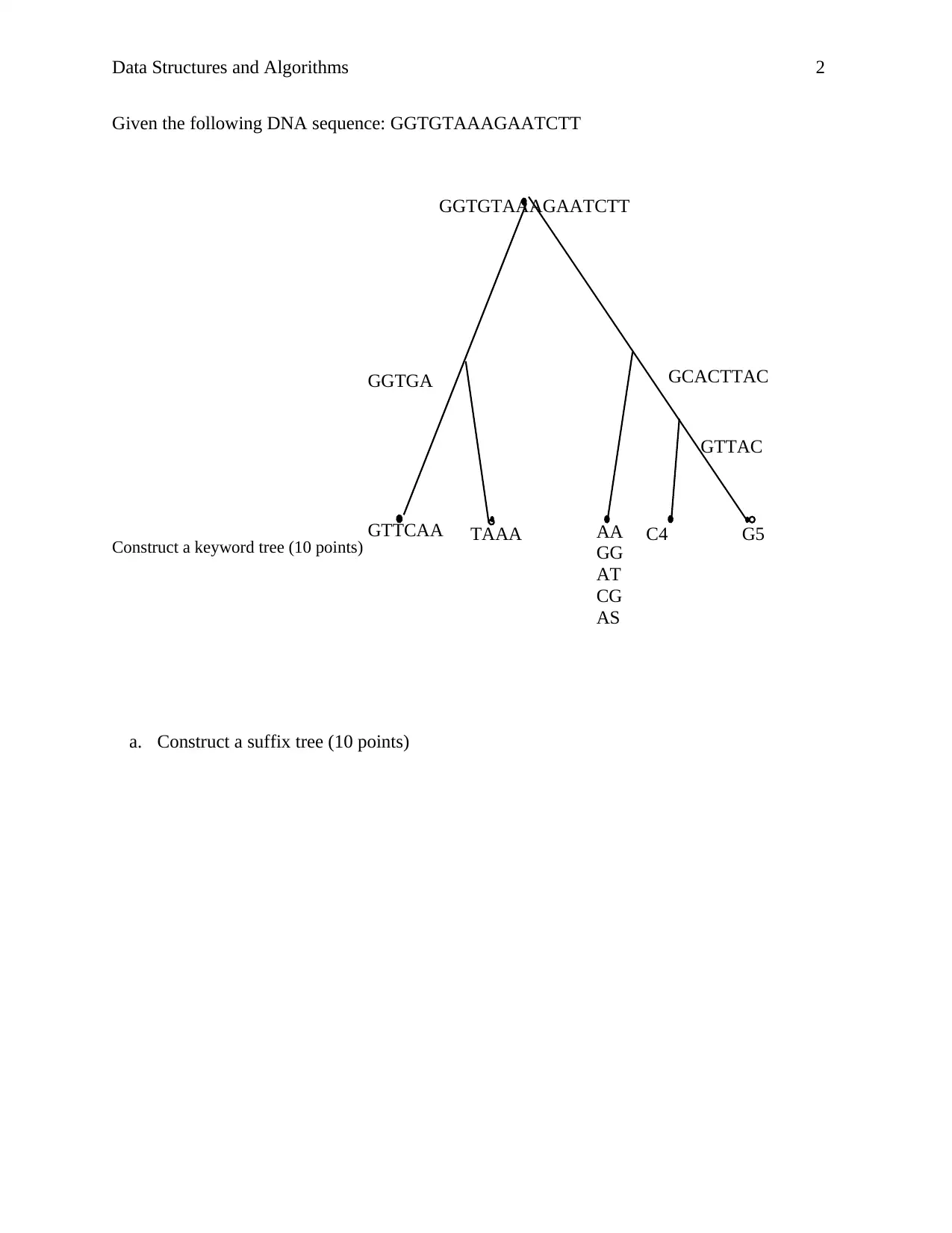
Given the following DNA sequence: GGTGTAAAGAATCTT
Construct a keyword tree (10 points)
a. Construct a suffix tree (10 points)
GTTCAA TAAA AA
GG
AT
CG
AS
C4 G5
GTTAC
GCACTTACGGTGA
GGTGTAAAGAATCTT
Given the following DNA sequence: GGTGTAAAGAATCTT
Construct a keyword tree (10 points)
a. Construct a suffix tree (10 points)
GTTCAA TAAA AA
GG
AT
CG
AS
C4 G5
GTTAC
GCACTTACGGTGA
GGTGTAAAGAATCTT
Data Structures and Algorithms 3
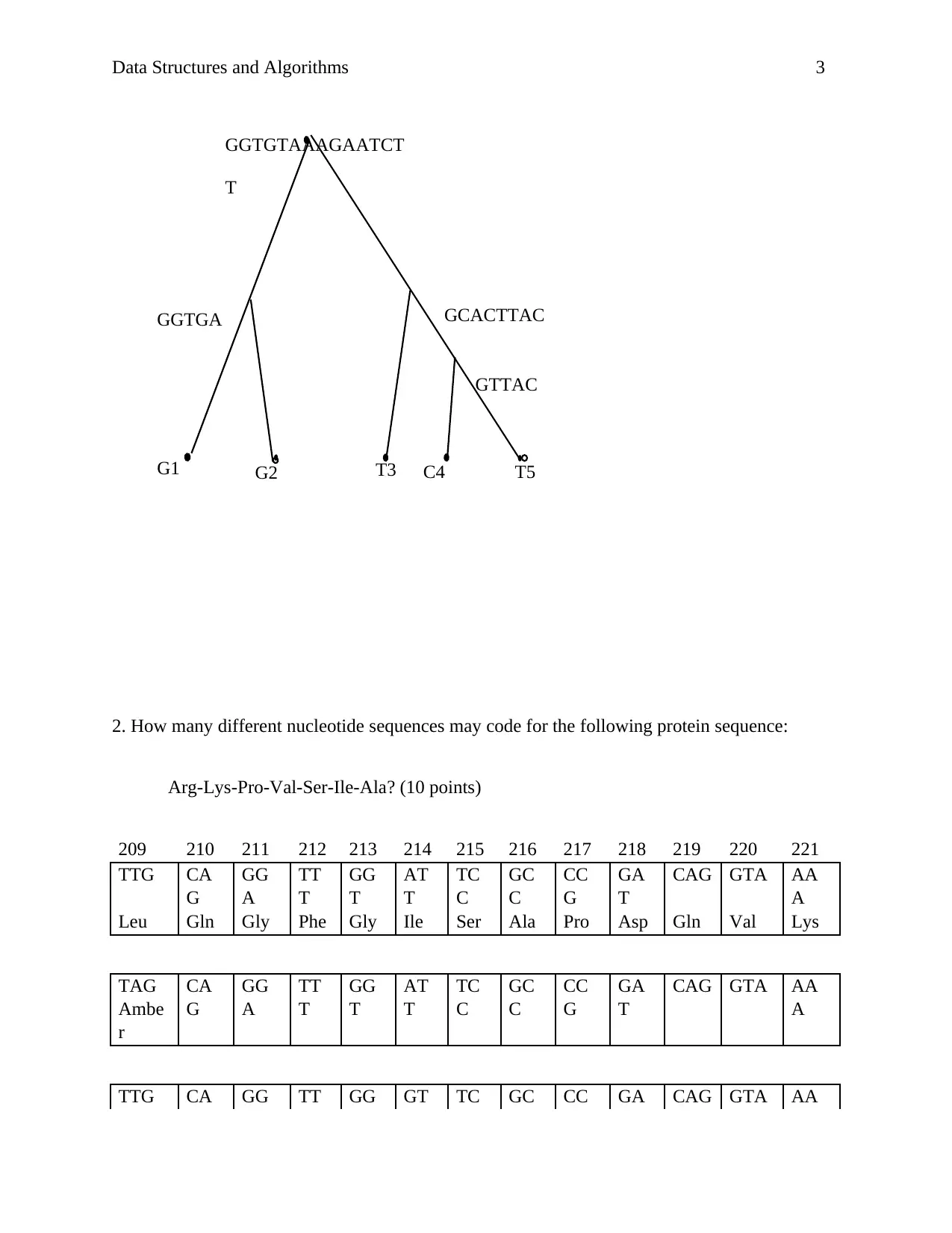
2. How many different nucleotide sequences may code for the following protein sequence:
Arg-Lys-Pro-Val-Ser-Ile-Ala? (10 points)
209 210 211 212 213 214 215 216 217 218 219 220 221
TTG CA
G
GG
A
TT
T
GG
T
AT
T
TC
C
GC
C
CC
G
GA
T
CAG GTA AA
A
Leu Gln Gly Phe Gly Ile Ser Ala Pro Asp Gln Val Lys
TAG
Ambe
r
CA
G
GG
A
TT
T
GG
T
AT
T
TC
C
GC
C
CC
G
GA
T
CAG GTA AA
A
TTG CA GG TT GG GT TC GC CC GA CAG GTA AA
G1 G2 T3 C4 T5
GTTAC
GCACTTACGGTGA
GGTGTAAAGAATCT
T
2. How many different nucleotide sequences may code for the following protein sequence:
Arg-Lys-Pro-Val-Ser-Ile-Ala? (10 points)
209 210 211 212 213 214 215 216 217 218 219 220 221
TTG CA
G
GG
A
TT
T
GG
T
AT
T
TC
C
GC
C
CC
G
GA
T
CAG GTA AA
A
Leu Gln Gly Phe Gly Ile Ser Ala Pro Asp Gln Val Lys
TAG
Ambe
r
CA
G
GG
A
TT
T
GG
T
AT
T
TC
C
GC
C
CC
G
GA
T
CAG GTA AA
A
TTG CA GG TT GG GT TC GC CC GA CAG GTA AA
G1 G2 T3 C4 T5
GTTAC
GCACTTACGGTGA
GGTGTAAAGAATCT
T
⊘ This is a preview!⊘
Do you want full access?
Subscribe today to unlock all pages.

Trusted by 1+ million students worldwide

Data Structures and Algorithms 4
G A T T T C C G T A
Leu Gln Gly Phe Gly Val Ser Ala Pro Asp Gln Val Lys
∆
TTG CA
G
GG
A
TT
T
GG
T
AT
T
TC
C
GC
C
CC
G
ATC AG
G
TAA AA-
Leu Gln Gly Phe Gly Ile Ser Ala Pro Ile Arg Ochr
e
Therefore 52 different nucleotide sequences.
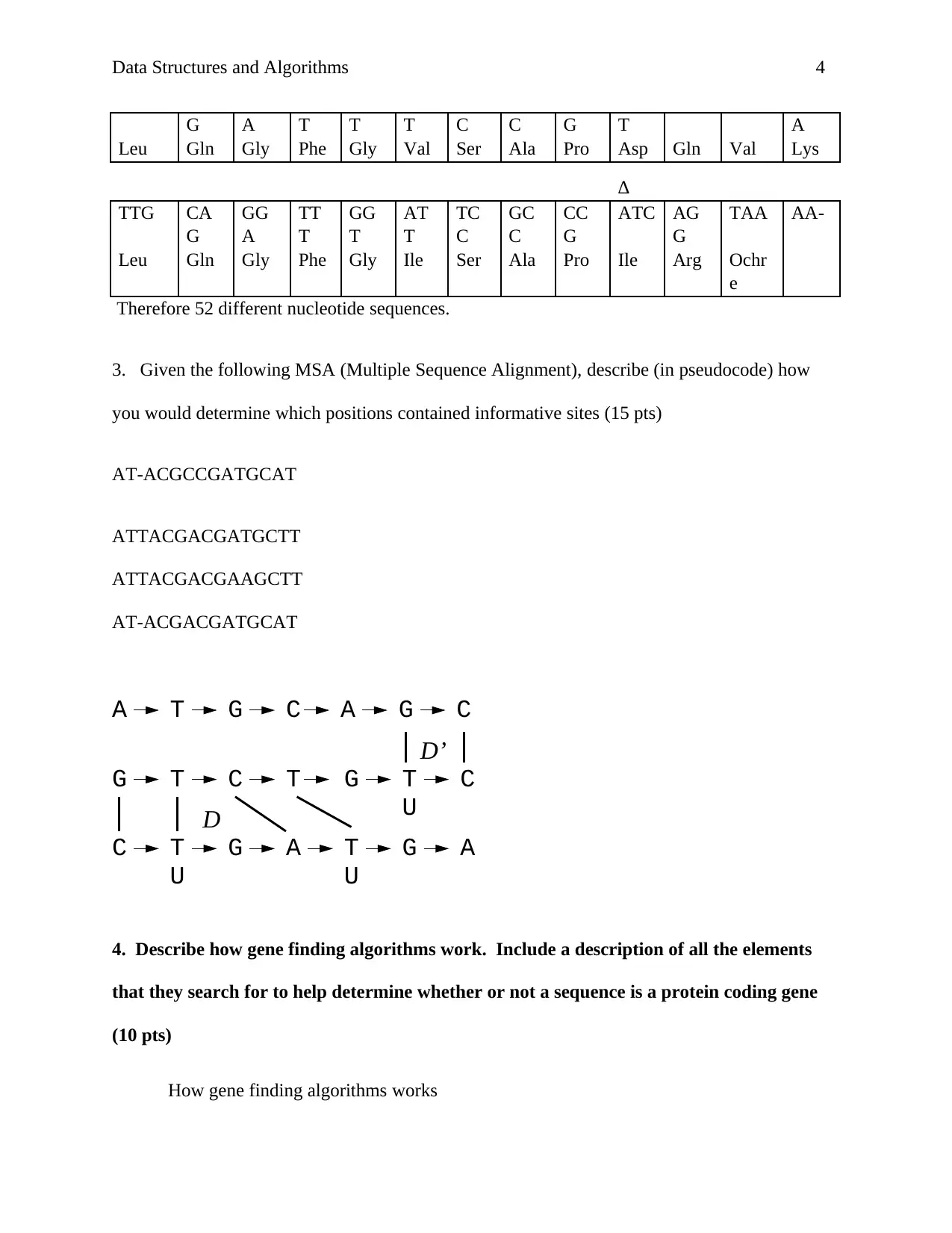
3. Given the following MSA (Multiple Sequence Alignment), describe (in pseudocode) how
you would determine which positions contained informative sites (15 pts)
AT-ACGCCGATGCAT
ATTACGACGATGCTT
ATTACGACGAAGCTT
AT-ACGACGATGCAT
4. Describe how gene finding algorithms work. Include a description of all the elements
that they search for to help determine whether or not a sequence is a protein coding gene
(10 pts)
How gene finding algorithms works
T G C
T CG
T
U
G A
A G C
T
U
G A
T
U
CG
A
T
C
D
D’
G A T T T C C G T A
Leu Gln Gly Phe Gly Val Ser Ala Pro Asp Gln Val Lys
∆
TTG CA
G
GG
A
TT
T
GG
T
AT
T
TC
C
GC
C
CC
G
ATC AG
G
TAA AA-
Leu Gln Gly Phe Gly Ile Ser Ala Pro Ile Arg Ochr
e
Therefore 52 different nucleotide sequences.
3. Given the following MSA (Multiple Sequence Alignment), describe (in pseudocode) how
you would determine which positions contained informative sites (15 pts)
AT-ACGCCGATGCAT
ATTACGACGATGCTT
ATTACGACGAAGCTT
AT-ACGACGATGCAT
4. Describe how gene finding algorithms work. Include a description of all the elements
that they search for to help determine whether or not a sequence is a protein coding gene
(10 pts)
How gene finding algorithms works
T G C
T CG
T
U
G A
A G C
T
U
G A
T
U
CG
A
T
C
D
D’
Paraphrase This Document
Need a fresh take? Get an instant paraphrase of this document with our AI Paraphraser

Data Structures and Algorithms 5
Since gene finding refers to the process of identification of the parts of the genomic DNA that
usually encodes the protein coding genes, several approaches are employed to enhance the
prediction. The algorithm employs two main methods in searching the elements. In similarity
evidence based gene finding systems, the target genome is usually searched in the sequence that
are same and similar extrinsic evidence. In a protein sequence, the member class of possible
coding the DNA is obtained by reversing the translation of the genetic code. Once the candidate
DNA sequence has been determined, the target genome is then searched that matches. The
matches can be partial, complete, exact or even inexact. Consequently, a high similarity degree
to a protein product gives a strong evidence that the region of the target genome is a protein
coding gene.
5. What is BLAST? Describe how the algorithm works. Be sure to include any statistical
measures that are used in determining the strength of any BLAST results. (10 pts)
Basic Local Alignment Search Tool (BLAST) in bioinformatics is a widely used program
and it refers to the algorithm used for comparing the primary biological sequence information
such the protein sequences, the DNA sequences, the amino acid sequences and even the
nucleotides.
How the BLAST algorithm works?
Normally to run the algorithm, the BLAST usually needs a query sequence to search
against (usually the target sequence) and the sequence to search for. Then the BLAST finds the
respective subsequences in the database which are also the same and similar to the sub sequences
in the query (Casey, p 22). In most usage, the query may be more than the database.
When the algorithm searches a sequence database, it first computes the pairwise
Since gene finding refers to the process of identification of the parts of the genomic DNA that
usually encodes the protein coding genes, several approaches are employed to enhance the
prediction. The algorithm employs two main methods in searching the elements. In similarity
evidence based gene finding systems, the target genome is usually searched in the sequence that
are same and similar extrinsic evidence. In a protein sequence, the member class of possible
coding the DNA is obtained by reversing the translation of the genetic code. Once the candidate
DNA sequence has been determined, the target genome is then searched that matches. The
matches can be partial, complete, exact or even inexact. Consequently, a high similarity degree
to a protein product gives a strong evidence that the region of the target genome is a protein
coding gene.
5. What is BLAST? Describe how the algorithm works. Be sure to include any statistical
measures that are used in determining the strength of any BLAST results. (10 pts)
Basic Local Alignment Search Tool (BLAST) in bioinformatics is a widely used program
and it refers to the algorithm used for comparing the primary biological sequence information
such the protein sequences, the DNA sequences, the amino acid sequences and even the
nucleotides.
How the BLAST algorithm works?
Normally to run the algorithm, the BLAST usually needs a query sequence to search
against (usually the target sequence) and the sequence to search for. Then the BLAST finds the
respective subsequences in the database which are also the same and similar to the sub sequences
in the query (Casey, p 22). In most usage, the query may be more than the database.
When the algorithm searches a sequence database, it first computes the pairwise

Data Structures and Algorithms 6
alignment of the sequence against all known sequences in the database. Then the best and
scoring significant homologs are deleted. The algorithm employs some statistical methods and
approaches that are used to determine the strength of the BLAST. The two main statistical
methods employed is: Assessing Alignment Significance.
With this method, random alignments are randomly generated and their scores computed
at first step. The mean and the standard deviation of the random scores are then computed. The
deviation of the actual random scores from the mean of the score is then computed. From this,
evaluation of the significance of the alignment is then done.
6. A graduate student has written part of an R script to perform an analysis. It is listed
below.
Describe what each line does by adding comment lines to it as appropriate (10 pts)
#-------------------------------------------------------
library(stats) (refers to the statistical data set in a library)
mydata <- iris (data set to be executed)
# Round 1 ( refers to frequency of the execution)
set.seed(101) (means there are 101 results to be executed)
km <- kmeans(mydata[,1:4], 10) (10 refers to the number of centroids to be displayed
plot(mydata[,1], mydata[,2], col=km$cluster) ( refers to the axes names/labels on the
script)
points(km$centers[,c(1,2)], col=1:3, pch=19, cex=2) (refers to points at which the
alignment of the sequence against all known sequences in the database. Then the best and
scoring significant homologs are deleted. The algorithm employs some statistical methods and
approaches that are used to determine the strength of the BLAST. The two main statistical
methods employed is: Assessing Alignment Significance.
With this method, random alignments are randomly generated and their scores computed
at first step. The mean and the standard deviation of the random scores are then computed. The
deviation of the actual random scores from the mean of the score is then computed. From this,
evaluation of the significance of the alignment is then done.
6. A graduate student has written part of an R script to perform an analysis. It is listed
below.
Describe what each line does by adding comment lines to it as appropriate (10 pts)
#-------------------------------------------------------
library(stats) (refers to the statistical data set in a library)
mydata <- iris (data set to be executed)
# Round 1 ( refers to frequency of the execution)
set.seed(101) (means there are 101 results to be executed)
km <- kmeans(mydata[,1:4], 10) (10 refers to the number of centroids to be displayed
plot(mydata[,1], mydata[,2], col=km$cluster) ( refers to the axes names/labels on the
script)
points(km$centers[,c(1,2)], col=1:3, pch=19, cex=2) (refers to points at which the
⊘ This is a preview!⊘
Do you want full access?
Subscribe today to unlock all pages.

Trusted by 1+ million students worldwide
Data Structures and Algorithms 7
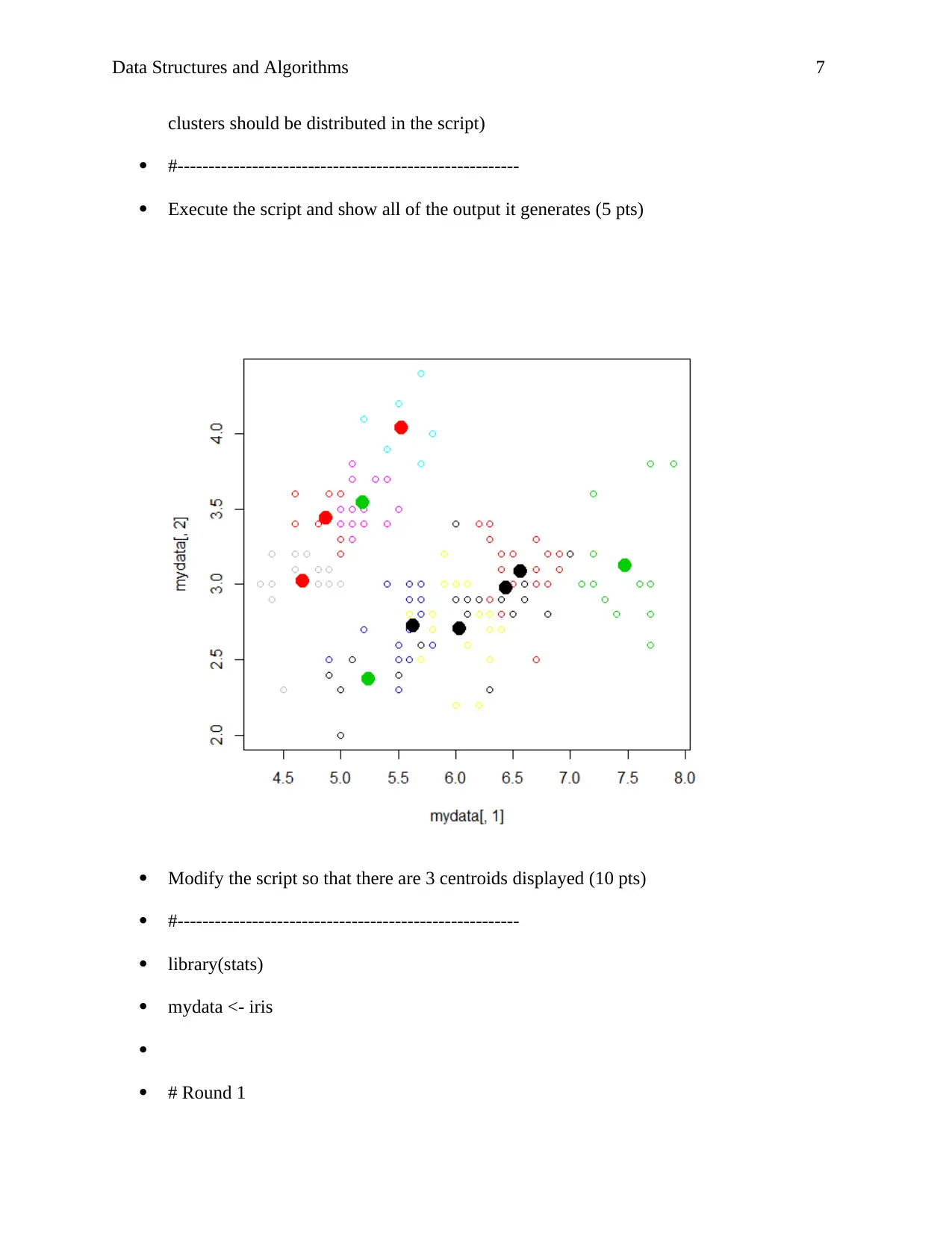
clusters should be distributed in the script)
#-------------------------------------------------------
Execute the script and show all of the output it generates (5 pts)
Modify the script so that there are 3 centroids displayed (10 pts)
#-------------------------------------------------------
library(stats)
mydata <- iris
# Round 1
clusters should be distributed in the script)
#-------------------------------------------------------
Execute the script and show all of the output it generates (5 pts)
Modify the script so that there are 3 centroids displayed (10 pts)
#-------------------------------------------------------
library(stats)
mydata <- iris
# Round 1
Paraphrase This Document
Need a fresh take? Get an instant paraphrase of this document with our AI Paraphraser
Data Structures and Algorithms 8
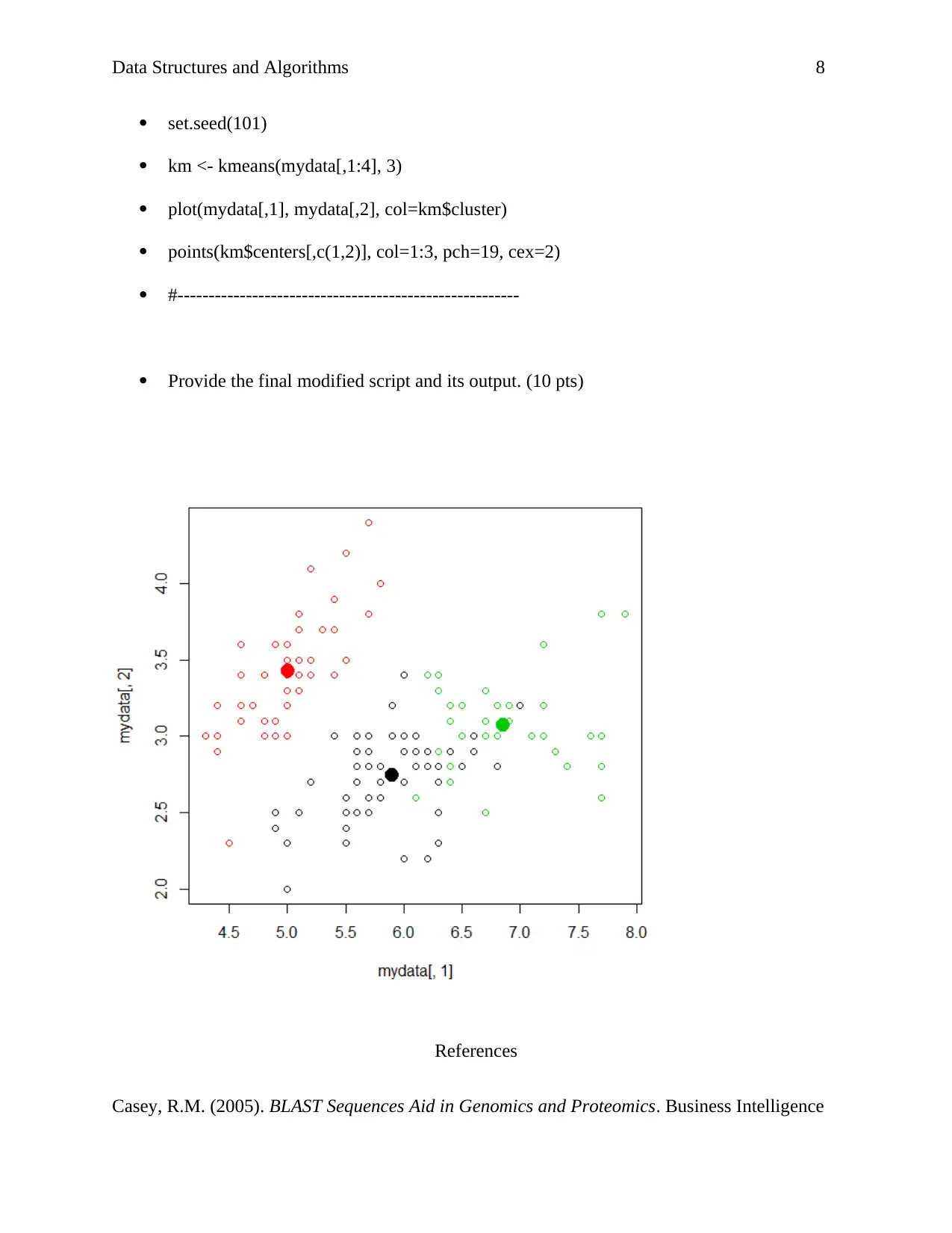
set.seed(101)
km <- kmeans(mydata[,1:4], 3)
plot(mydata[,1], mydata[,2], col=km$cluster)
points(km$centers[,c(1,2)], col=1:3, pch=19, cex=2)
#-------------------------------------------------------
Provide the final modified script and its output. (10 pts)
References
Casey, R.M. (2005). BLAST Sequences Aid in Genomics and Proteomics. Business Intelligence
set.seed(101)
km <- kmeans(mydata[,1:4], 3)
plot(mydata[,1], mydata[,2], col=km$cluster)
points(km$centers[,c(1,2)], col=1:3, pch=19, cex=2)
#-------------------------------------------------------
Provide the final modified script and its output. (10 pts)
References
Casey, R.M. (2005). BLAST Sequences Aid in Genomics and Proteomics. Business Intelligence

Data Structures and Algorithms 9
Network.
Mount, D. W. (2004). Bioinformatics Sequence and Genome Analysis. 2nd ed. Cold Spring
Harbor Press.
Network.
Mount, D. W. (2004). Bioinformatics Sequence and Genome Analysis. 2nd ed. Cold Spring
Harbor Press.
⊘ This is a preview!⊘
Do you want full access?
Subscribe today to unlock all pages.

Trusted by 1+ million students worldwide
1 out of 9
Related Documents
Your All-in-One AI-Powered Toolkit for Academic Success.
+13062052269
info@desklib.com
Available 24*7 on WhatsApp / Email
![[object Object]](/_next/static/media/star-bottom.7253800d.svg)
Unlock your academic potential
Copyright © 2020–2026 A2Z Services. All Rights Reserved. Developed and managed by ZUCOL.



